Learn About the History of Recombinant DNA Technology
Restriction Using Bacteriophages
While most freezers in laboratories are stocked with DNA sequencing material such as restrictive enzymes today, there was a time when this was not true. The history of recombinant DNA technology was a quest to find restrictive enzymes that could be used successfully to cleave DNA . The ability to utilize the concept of restriction was developed in the early 1950s. It was originally termed as host-induced variation. Bacteriophages placed within a certain host strain were able to grow. Once this growth was established, however, the phages would only continue to grow in the same strain. This influenced a generation of scientists to understand the properties of this growth.
Challenges with Restrictive Enzymes
During the early years of the 1960s, researchers struggled with identifying a restrictive agent that would cleave genes in a controlled manner. The majority of the enzymes could be used to recognize specific sequences of DNA, they would cleave the subject in random ways. This made them ineffective for cloning or reagent mapping.
Restrictive Endonuclease
In 1971, two researchers from Johns Hopkins University, Kathleen Danna and Daniel Nathans, identified usable restrictive enzymes, specifically endonuclease R, able to isolate fragments of simian virus 40 in DNA using electrophoresis and restriction enzymes. Despite this breakthrough, the fundamental understanding of the ultimate ramifications was not known at the time. However, Nathans later found that by cleaving the genetic material apart, it provided a way to map the virus as well as the exact origin of replication. This created a way to map DNA of nearly anything and created the basics of modern molecular biology. Nathans eventually joined Werner Arber and Hamilton Smith in accepting the 1978 Nobel Prize in Medicine.
Creating the Process Used in Recombinant DNA Technology
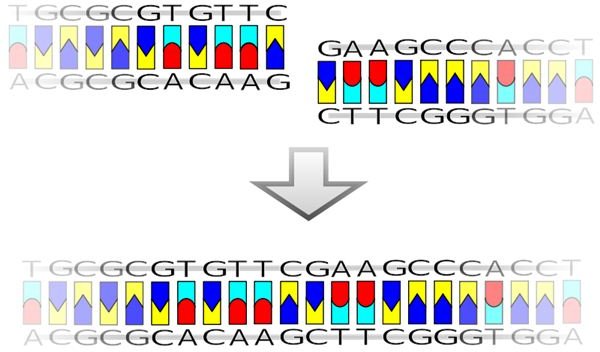
The technique that is used today as recombinant DNA was developed at the Department of Biochemistry at Stanford University by Peter Lobban and A. Dale Kaiser. In a series of papers published from 1972 to 1974, they described the process of isolating genes with restriction endonucleases and inserted them into plasmids. These plasmids were used as the vector for the process. They could then themselves be inserted into the target bacteria. This made the subject bacteria posses the DNA of a different organism because the plasmid can reproduce without the need for chromosomal DNA. Other famous scientists who also helped develop this process were Stanley Norman Cohen and Herbert Boyer, who were eventually awarded the patent in 1980 for the process to create functional molecular chimeras in bacteria that did not exist in nature. This brought the history of recombinant DNA technology to the modern age.
References
“How Restrictive Enzymes Became the Workhorses of Molecular Biology” Proceedings of the National Academy of Sciences: https://www.pnas.org/content/102/17/5905.full
“Genetics and Genomics Timeline” Genome News Network: https://www.genomenewsnetwork.org/resources/timeline/1973_Boyer.php
Image Source
Blunt Ligation. (Supplied by Madprime at Wikimedia Commons; GNU Free Documentation License; https://upload.wikimedia.org/wikipedia/commons/d/de/Blunt_ligation.svg)
This post is part of the series: The Genetics of Recombinant DNA
Recombinant DNA is the process of merging genes from one organism with the genes of another organism. The most common method to complete the process involves using restriction enzymes to isolate a gene and then place it on a vector plasmid. It is then inserted into a host cell such as bacteria.